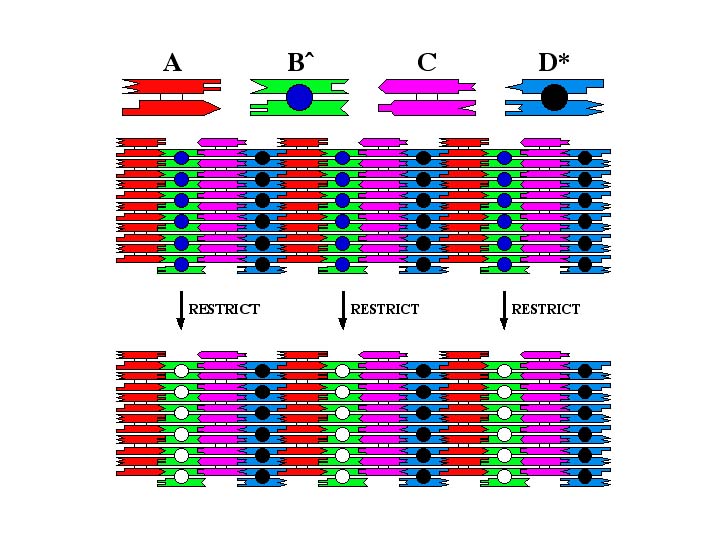

At the top of this drawing are four double crossover molecules, A B÷, C and D*, which are shown schematically. The pattern results from hairpins directed out of the plane of the array from molecules B÷ and D*. Hairpin B÷ contains a restriction site missing on D*. When the array is treated with the restriction enzyme, this hairpin is removed. Thus, the 32 nm pattern on the top is converted to a 64 nm pattern after restriction.
The second modificaton entails performing the reverse operation, as shown below:
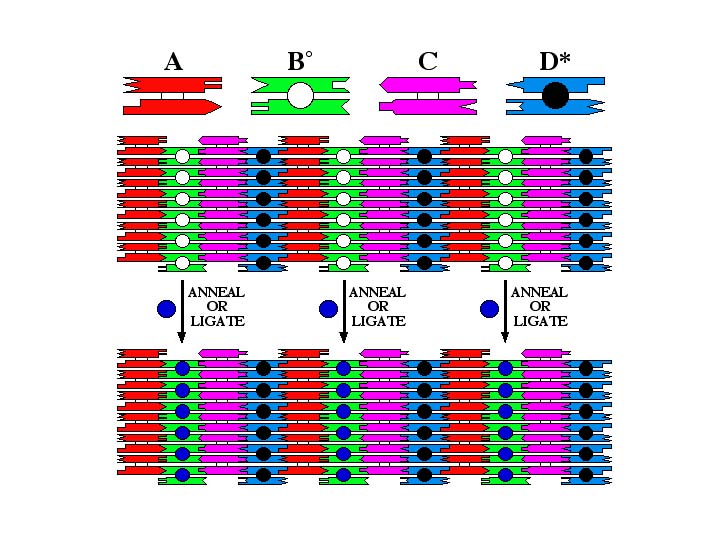
The components of the array here are similar to those above. However, we have replaced B÷ with a different component double crossover tile, B¹. This tile contains a sticky end, that can pair with a sticky end on a hairpin in solution. Thus, the upper array produces a 64 nm pattern, but when a hairpin is ligated to it, or even just hydrogen bonded to it, a 32 nm pattern results, as shown in the bottom array.
Thus, we can modify specific structural features created by self-assembly techniques.
Return to 2-D DNA crystals.
Return to Ned Seeman's Nanotechnology Page
Return to Homepage for Ned Seeman's Lab